 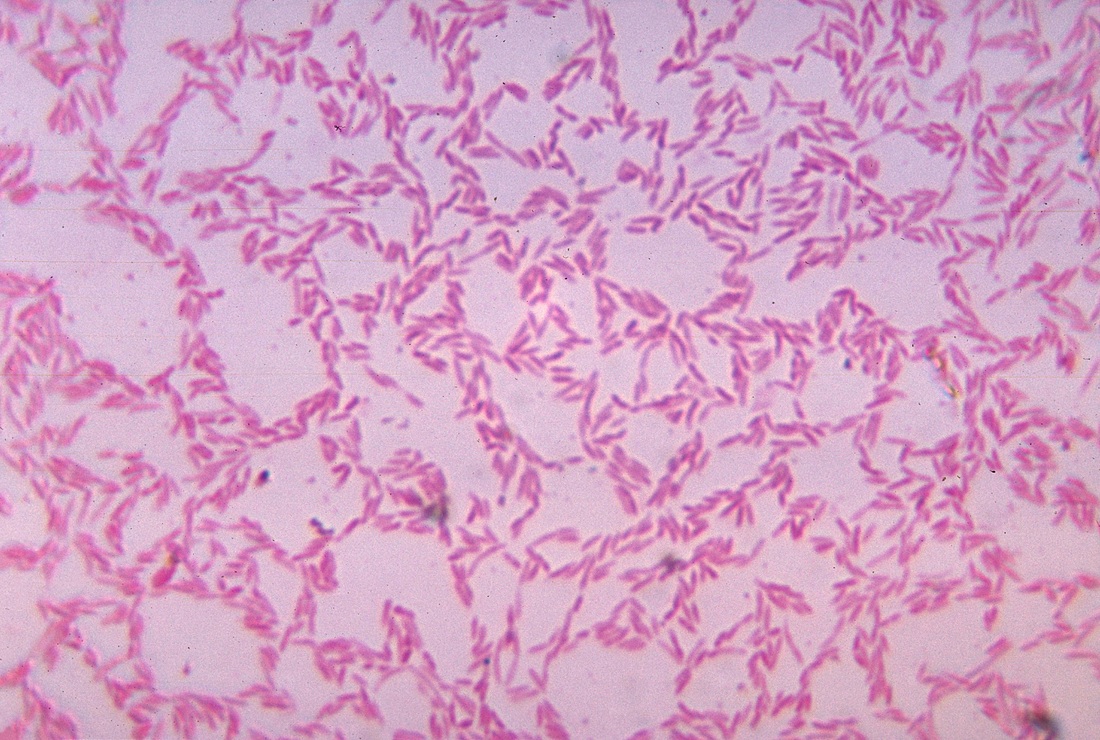
coli are responsible for various clinical manifestations ranging from diarrhoea, urinary tract infections, and life-threatening septicemia, resulting in over 2 million deaths every year globally ( Hussain et al., 2012 Huang et al., 2021). However, infections with variant strains of E. 
Its primary habitat includes the lower intestinal tract of humans and animals ( Nia Santos Braz et al., 2020). It is the most common organism responsible for opportunistic infections. coli that would help in better treatment and prevention strategies.Įscherichia coli is a versatile gram-negative bacterium with huge genetic diversity ( Nia Santos Braz et al., 2020). Further, there is a need to undertake systematic, unbiased monitoring of predominant lineages amongst non-lactose fermenting E. coli be considered equally capable as lactose fermenting forms in causing intestinal and extraintestinal infections. We suggest that non-lactose fermenting E. Our study underscores the presence of critical pathogens and MDR clones amongst non-lactose fermenting E. coli pathotypes demonstrated β-hemolysis. The bla CTX-M-15 extended-spectrum beta-lactamase gene was present in 53% of isolates (9/17), whilst 64.7% (11/17) isolates were affiliated with pathogenic pathotypes. The predominant phylogroup detected was B2 (65%). Phylogenetic analysis corroborated a striking clonal population amongst the studied NLF E. WGS data analysis revealed international high-risk clonal lineages. coli and could not ferment lactose sugar. All isolates (17/17) were confirmed as E. coli was 10%, of which 47% (8/17) exhibited multi-drug resistant (MDR) phenotypes. coli from a diagnostic centre (icddr,b) in Dhaka, Bangladesh. To address this gap, we used whole-genome sequencing (WGS) coupled with detailed microbiological and biochemical testing to investigate 17 NLF E. Although several studies have characterised such strains using conventional methods, they have not been comprehensively studied at the genomic level. coli) are responsible for various intestinal and extraintestinal infections. Multi-resistant pathogenic strains of non-lactose fermenting Escherichia coli (NLF E. 4Biosafety and BS元 Laboratory, Biosafety Office, International Centre for Diarrhoeal Disease Research, Bangladesh (icddr,b), Dhaka, Bangladesh.3Clinical Microbiology and Immunology Laboratory, Laboratory Sciences and Services Division, International Centre for Diarrhoeal Disease Research, Bangladesh (icddr,b), Dhaka, Bangladesh.2Department of Infection Biology, London School of Hygiene and Tropical Medicine, London, United Kingdom.1Laboratory Sciences and Services Division, International Centre for Diarrhoeal Disease Research, Bangladesh (icddr,b), Dhaka, Bangladesh.Arefeen Haider 1 Araf Mahmud 1 Dilruba Ahmed 3 Md Asadulghani 4 Taane G. Razib Mazumder 1 † Arif Hussain 1 † Jody E. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
